Doose Syndrome. In the early 1970s, a group of children with severe childhood epilepsies was found to have comparable clinical features that consisted of quick jerks and subsequent drop attacks amongst other types of epileptic seizures. These seizures, myoclonic-astatic or myoclonic-atonic seizures, eventually became the defining feature of an epilepsy syndrome referred to as Myoclonic Astatic Epilepsy or Doose Syndrome. In the recent issue of the American Journal of Human Genetics, we report on the first true gene for Doose Syndrome. Here is the story of SLC6A1 (GAT-1).
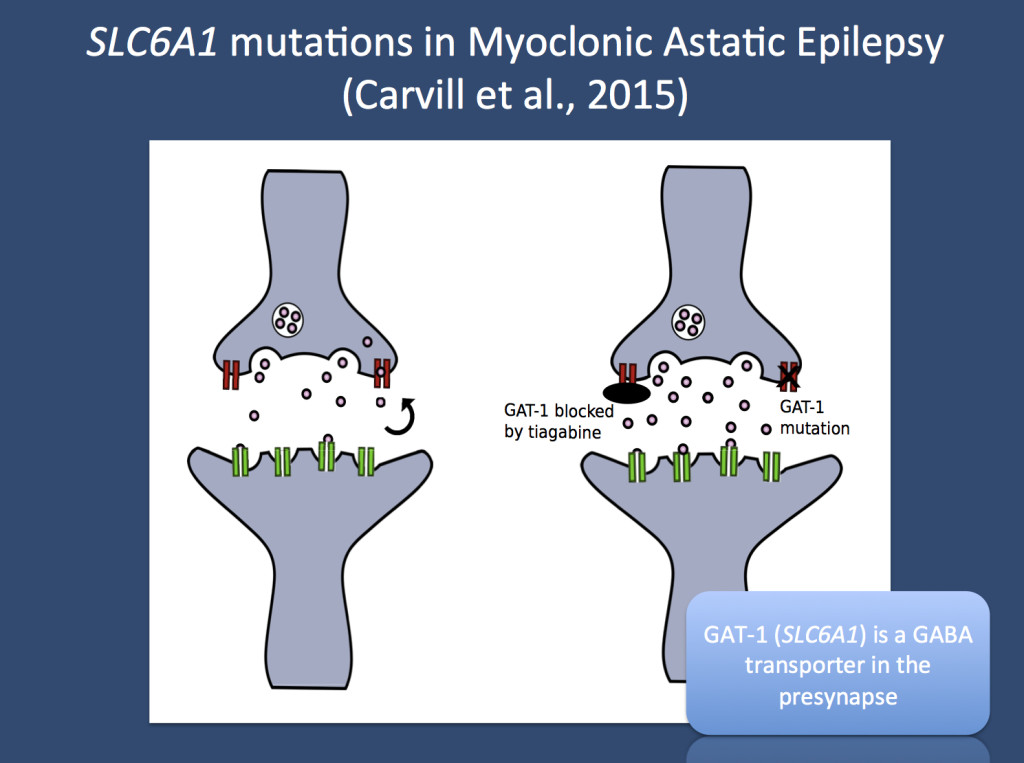
GAT-1 transports GABA back into the presynaptic terminal. When GAT-1 is blocked by tiagabine or if GAT-1 activity is reduced by mutations in the SLC6A1 gene, GAT-1 transport is impaired and GABA accumulates in the synapse. Pink dots – GABA, red – GAT-1, green – GABA receptor. (Image provided by Gemma Carvill and Heather Mefford, Seattle)
Travel post part 2. I was initially prepared to write my second blog post on our vacation in Europe this week and was initially planning on featuring our time in Kiel. However, then our publication on SLC6A1 came out. As I have always struggled with blogging about conferences or our research travel, I thought that it would be appropriate to honor our time in Kiel by writing about a prominent gene finding in the epilepsy syndrome that is intrinsically connected to this city in Northern Germany and was named after the former head of child neurology in Kiel: Doose Syndrome (BTW the correct pronunciation is “Dohs-ah” and doesn’t rhyme with “goose”). It took more than 40 years from conceptualizing this entity to identifying the first genuine gene for this syndrome, which is now reported in the recent study by Carvill and collaborators. We were involved in this project through the follow-up projects of the EuroEPINOMICS RES working groups.
Doose Syndrome. There is a group of childhood epilepsies that has always been very hard to conceptualize. This group includes children with severe epilepsies and multiple seizure types, various degrees of cognitive impairment, variable outcomes and a complex, often refractory response to antiepileptic medication. Prior to the advent of high-resolution neuroimaging and genetic testing, there were few objective ways of capturing a correlate of the epilepsy, and many clinicians turned to the EEG as a tool for predicting and classifying epilepsies. However, even when children with epilepsy and a generalized EEG and normal neuroimaging were grouped together, finding the right boundary to delineate syndromes was challenging. Or, as Doose himself put it in 1992: “There is hardly another field in pediatric epileptology presenting such terminological uncertainty and confusion as is to be found in the domain of epileptic syndromes with generalized minor seizures of early childhood.” (H. Doose in Epileptic syndromes in infancy, childhood and adolescence, 2nd Edition 1992). However, one group of patients stood out in particular: children with myoclonic-astatic/atonic seizures and a generalized EEG pattern. This group was initially called “centrencephalic myoclonic-astatic petit mal”, which eventually became Myoclonic Astatic Epilepsy (MAE) or Doose Syndrome. Given the lack of identifiable causative factors, a genetic predisposition was assumed early on. Some initial studies looked at the inheritance of the EEG features and suggested mainly a familial predisposition. It turns out that these initial concepts were both right and wrong: Doose Syndrome has become an important, recognizable clinical entity, but the genetic predisposition may rather be due to de novo mutations than to familial factors.
SLC6A1. In the recent publication by Carvill and collaborators, we screened a large group of 644 patients with epileptic encephalopathies for mutations in SLC6A1, a gene that was initially identified in one of the large exome studies. When the phenotype of the initial patient reported by Rauch and collaborators was investigated in detail, it turned out to be a patient with MAE. Given that other genetic causes for MAE are only rarely found and usually represent genes that are also found in other neurodevelopmental diseases, this was an interesting starting point. SCL6A1 mutations were subsequently found in 7 patients with epileptic encephalopathies, and most of these mutations were de novo – interestingly all patients had an MAE phenotype, accounting for almost 5% of patients with MAE. Two mutations were truncating mutations, suggesting that the disease mechanism is haploinsufficiency, the lack of one of the two gene products.
The GABA recycler. SLC6A1, also called GAT-1, has an important role in the brain. The GAT-1 transporter removes GABA from the synaptic cleft. GABA is the main inhibitory transmitter in the brain, the main messenger that says “no” in the CNS. In order for proper functioning of neuronal networks, a balanced activity of the excitatory neurotransmitters such as glutamate and the inhibitory transmitters such as GABA is needed. If this balance is disturbed, epilepsy may occur. GAT-1 is no stranger to the field of epileptology as it represents the protein that the antiepileptic drug tiagabine (Gabitril) binds to. Herein lies the problem in understanding how SLC6A1 mutations cause epilepsy: SLC6A1 mutations reduce the functional amount of GAT-1 on the presynapse (figure), which should actually increase both the duration and the quantity of GABA in the synaptic cleft. This should give us more GABA rather than too little. How the supposedly increase in GABA will lead to epilepsy is entirely unclear – however, it provides us with a good example of why we might sometimes be safer looking at genetic data on a larger scale rather than relying on our instincts regarding gene function.
This is what you need to know. Even though individual patients with MAE have an established genetic cause such as SLC2A1 mutations, microdeletions or novel epilepsy genes, the majority of MAE remains to be explained. With SCL6A1, we have a new gene, which may explain up to 5% of all MAE patients. Therefore, SLC6A1 is the first proper gene for Doose Syndrome. However, let me back pedal a little. Calling SLC6A1 is “the Doose gene” is probably overstated. Doose Syndrome is likely to be genetically heterogeneous and – in contrast to Dravet Syndrome – caused by various different genes. It will be interesting to see progress in this area in the near future.



Great news that some progress is being made to understand this disease. Thanks for posting this fantastic news.