EIEE. Online Mendelian Inheritance in Man (OMIM) is the undisputed main resource for information regarding genes and disease. It is the resource that the majority of clinicians and researchers in the field turn to in order to get information about established or novel genetic etiologies in genetic epilepsies and neurodevelopmental disorders. However, historically, OMIM had decided to enumerate many of the genes for developmental and epileptic encephalopathies within a phenotypic series called Early Infantile Epileptic Encephalopathies (EIEE). The field has advanced, and we now understand that most genetic etiologies have a broad phenotypic range and can cause a wide range of epilepsy phenotypes. Accordingly, in collaboration and consultation with our ClinGen epilepsy clinical domain working group, OMIM will retire the EIEE classification and refer to them as developmental and epileptic encephalopathies (DEE). Dravet Syndrome, formerly EIEE6 will now become DEE6, which is the secondary annotation to the actual term “Dravet Syndrome”. For some, this might be a small change in semantics. However, as a clinician trying to make sure that the uniqueness and distinctiveness of childhood epilepsies in the era of large-scale data analysis is appreciated, this small step is likely to be highly influential in the future. Here is some background on how the EIEEs finally became DEEs.
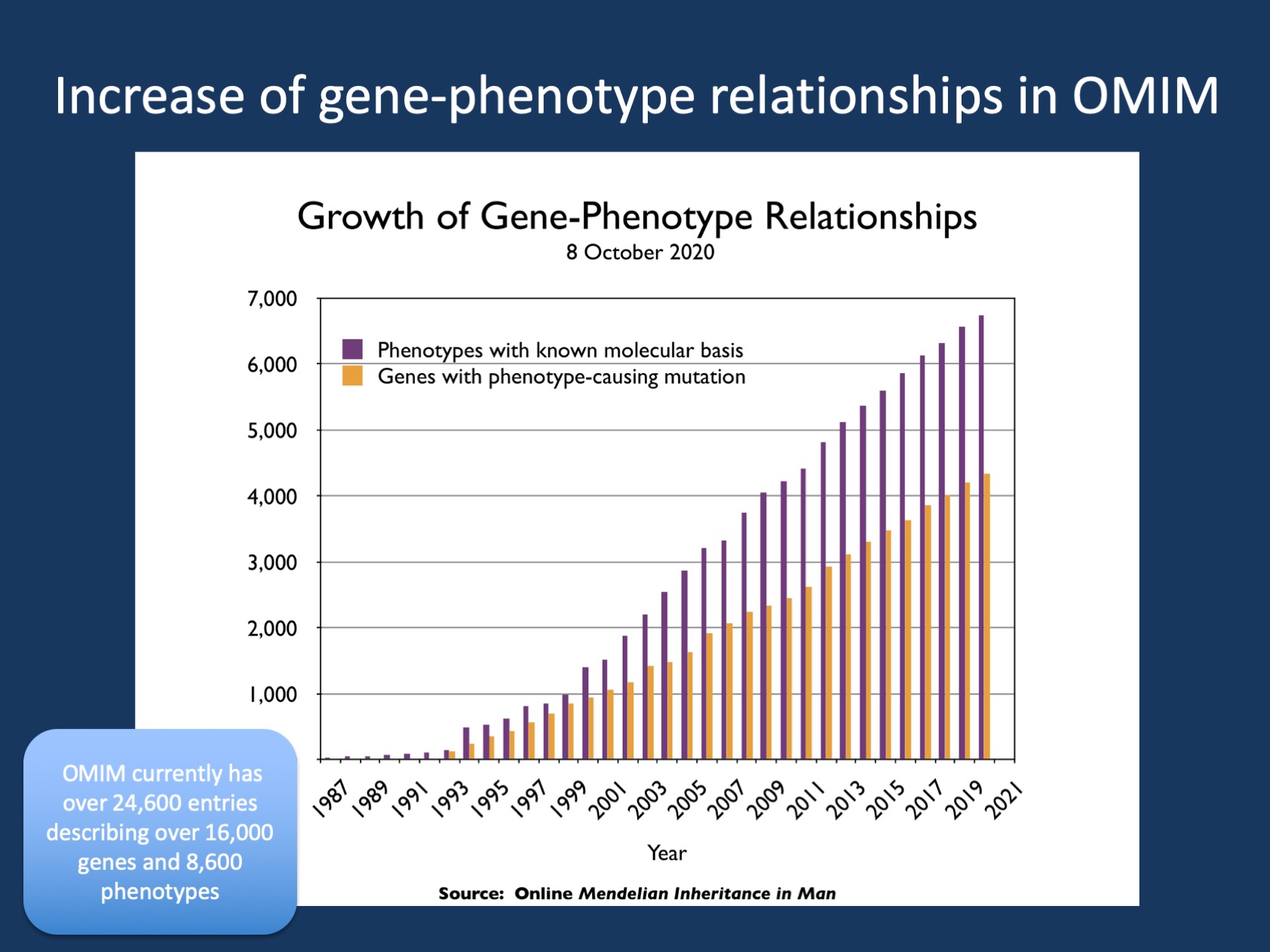
Figure 1. The pace of disease gene discovery as cataloged by the OMIM Morbid Map Scorecard. The current scorecard is available from https://omim.org/statistics/geneMap. The figure is an update provided by OMIM from the figure in Amberger et al., 2018.
OMIM. Without attempting to be repetitive, if you come across a gene you have not heard of, you will likely turn to OMIM as your resource. OMIM has over 24,600 entries describing over 16,000 genes and 8,600 phenotypes and represents a continuation of Mendelian Inheritance in Man (MIM) that initially started in 1966 by Victor A. McKusick. OMIM is one of the few trusted online resources that has been continuously maintained and updated. For the epilepsies, OMIM uses several so-called phenotypic series and the “early epileptic encephalopathy (EIEE)” series is the largest epilepsy-related curated list. Currently, this list ranges from EIEE1 (ARX) to EIEE88 (MDH1), a recently identified recessive developmental and epileptic encephalopathy characterized by global developmental delay, epilepsy, and progressive microcephaly. In addition to clinical information, this list of genes is also used for bioinformatic approaches, annotations, and interpretation of genomic information. Having written a blog on epilepsy genetics for the last decade, I am very much aware of the needs of staying updated with information. OMIM has consistently done this. However, some naming conventions have been historical. Hence, we have had our eyes on the EIEEs for quite a while – now, we finally get to address them!
ClinGen. I am part of the ClinGen Epilepsy Clinical Domain Working Group (CDWG) – when people ask me what ClinGen is, I typically answer that we define epilepsy genes. ClinGen is a National Institutes of Health (NIH)-funded resource to build a central resource that defines the clinical relevance of genes and variants for use in precision medicine and research. I don’t mean to boast, but there are few institutions and mechanisms to critically review and determine whether genes that are suggested in the literature are truly disease-causing and to what clinical features they are linked. The literature alone cannot generate this. With the focus on novelty in the scientific literature and the lack of incentive to report negative findings, “retiring” genes has become one of our tasks – think of EFHC1, SCN9A, RELN, ICK, CACNA1H – if you find these genes on a gene panel or exome report, this is against the current official categorization that you can find in the ClinGen database. These genes are not genetic etiologies for the epilepsies.
Retiring the EIEEs. When you curate epilepsy genes according to ClinGen criteria for 4+ years, you tend to run into issues how gene-disease relationships are typically described in common databases. Basically, labelling all severe epilepsies starting in the first year of life as EIEE creates difficulties. “Early Infantile” refers to the first three months of life when conditions like Dravet Syndrome (SCN1A) or Infantile Spasms (DNM1) typically don’t start yet. Moreover, nearly all genetic epilepsies are characterized by phenotypic variability and using EIEE for conditions like SLC6A1 or SYNGAP1 simply does not feel right. Doose Syndrome (Myoclonic Astatic Epilepsy) is a recurrent presentation of these genetic epilepsies, and a significant number of individuals never have seizures. Labelling these conditions as EIEE may suggest that they are more homogeneous than they actually are with respect to epilepsy phenotypes and does not take into account that most conditions also have other prominent neurological features such as intellectual disability, autism, or movement disorders. Therefore, as of August 2020, OMIM has agreed to successively change the EIEE terminology to the generally accepted language in the epilepsy field – developmental and epileptic encephalopathies.
What you need to know. OMIM has agreed to change the EIEE phenotypic series, renaming them to the generally accepted DEE terminology. This may not be a big step, but it highlights something important. The language we use in clinical epileptology on one hand and data curation, laboratory science, and data-driven approaches have an inherent tendency to drift apart. It is our task to re-connect these domains whenever possible to make sure that the language we use clinically to describe the complex presentations of individuals with DEE is also reflected in the way that the associated genetic causes are catalogued. Therefore, while renaming the EIEEs is only a small step, it emphasizes something much bigger – the active involvement of the epilepsy community in making sure that the complexity of the epilepsies is not left behind.


