rs6732655. I acknowledge that the title of this blog post looks like my keyboard is broken, but please bear with me. Last month, I blogged about a recent genome-wide association by the BioBank Japan (BBJ), discussing the evidence for a Single Nucleotide Polymorphism (SNP) in the vicinity of the SCN1A gene (rs6732655). In a prior study, the SNP in question was initially found to be associated with epilepsy and I discussed the fact that this SNP, albeit not significant by itself, was also seen at a higher frequency in cases than in controls in the epilepsy cohort of the BBJ study. I received some comments regarding this post and it was pointed out that my reasoning was incorrect given that rs6732655 was not nominally significant in the BBJ study. Therefore, this study was not a replication study in itself. Let me retrace my steps and revisit where my hunch came from to write the initial blog post.
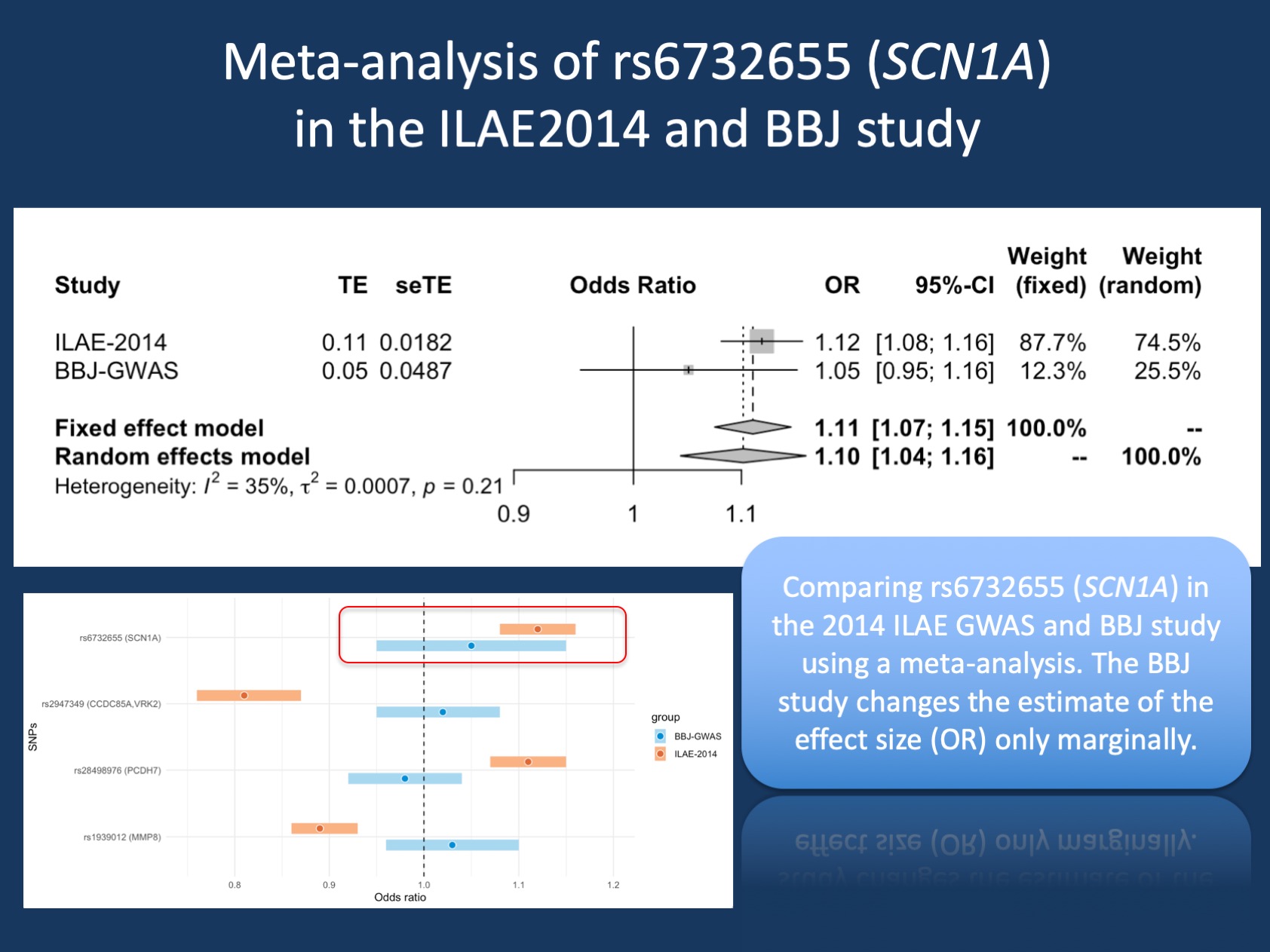
Figure 1. A meta-analysis of rs6732655 (SCN1A) in both studies mentioned in my prior blog post. The diamond-shaped symbol at the bottom of the Forest plot describes the overall estimated effect sizes using two different models, which basically result in the same estimate. Overall, the information of the BBJ study changes the estimate of the effect size (OR) only marginally. The joint estimate from both studies still suggests an OR of 1.1.
The falsification principle. When Austrian-British philosopher Karl Popper initially came up with the falsification principle, people didn’t realize how this concept would affect the way we make sense of information around us, especially within the context of scientific questions. Popper was concerned about distinguishing science from pseudo-science, and he drew the distinction between hypotheses we can disprove and hypotheses that cannot be disproven. Science works by putting forward a hypothesis and, while we can never prove our hypothesis, just a single example is sufficient to disprove it – take “all swans are white” as an example. We can use this principle for hypothesis testing in genetics. If we would like to find support for the hypothesis that SCN1A variants are associated with common epilepsies (yes, I am coming back to the topic of this post), then we can generate evidence for this by hypothesizing the opposite and find a good reason why this hypothesis, called the null hypothesis, cannot be true.
The degree of surprise. The degree of our surprise is reflected in the p-value. If we hypothesize that a given SCN1A variant (rs6732655) is not associated with epilepsy and we find that we reject this null hypothesis with p=0.05, it means that the observed distribution of rs6732655 in cases and controls would be expected in 1:20 random scenarios (or less). In other words, let’s assume that the frequency of rs6732655 in cases and controls is the same and see what we expect if we randomly chose a group of individuals. We get a p-value of 0.05 if the true frequency in cases of controls is so extreme that it would occur in 1:20 situations or less. We then reject our null hypothesis and while we can never prove that rs6732655 is truly associated, we can repeatedly DIS-prove that it is not. However, there is still a 5% chance that rs6732655 has nothing to do with epilepsy when we approach it this way and take the p-value at face value.
BBJ. To state it bluntly, rs6732655 is not significantly associated with epilepsy in the BBJ study. Therefore, this study does not confirm this marker is associated with epilepsy, and no conclusion can be drawn from this study as an independent replication. We might speculate that this variant may have been more prominent with better phenotyping (the BBJ cohort was likely quite heterogeneous) or larger numbers, but this is speculative and not the point. In summary, using the agreed-upon criteria on how we interpret study results, the BBJ study does not independently replicate the association of rs6732655 and epilepsy, and there is nothing more to it. Let me stop here for a second.
Meta-analysis. What we have done so far is evaluate the evidence for this SNP based on relatively conservative statistical criteria. Thinking about statistical evidence in a very conservative way is relatively commonplace in the “omics” field. We can do so many tests in parallel that we have to account for this, given that we are always tempted to pick and choose that data that fits our ideas. Historically, the tendency to over-emphasize findings that were barely significant was one of the reasons that genetic association studies struggled with replication, giving rise to the concept such as genome-wide significance. However, given that the data used for our studies is usually released in discrete chunks (individual studies or publications), methods have been developed to look at data across studies, including methods for meta-analysis. Here, I have used the meta package to assess the joint effect size in both studies (Figure 1). Overall, the BBJ study does very little to the estimate of the rs6732655 effect size across both studies. It slightly drags the estimated OR down, but barely affects the estimate and the 95% confidence interval.
What you need to know. Here, I have retraced my earlier steps and have refined my prior statement regarding the associations of rs6732655 (SCN1A) and epilepsy. I had written my initial blog post based on the “surprise” that the SNP in vicinity of SCN1A was the only one of four variants pointing in the same direction as in the initial ILAE study. However, as discussed in this post, this “hunch” can be misleading as it makes us pick and choose variants that seem to fit a pattern that we would like to see. Methods such as meta-analysis provide a formal process to answer these questions. The somewhat sobering result of a first, preliminary analysis suggests that the “directionality” that initially motivated me to write this post was in fact deceiving. The inclusion of the BBJ results neither significantly changes the odds ratio nor the confidence interval. I chose the comparison of ILAE 2014 vs. BBJ as the association of common variants in the vicinity of SCN1A and epilepsy is well-established in 2020. Several other studies, such as the large 2018 ILAE GWAS have supported such an association since the initial discovery.


