Genetic literacy. Sometimes important milestones don’t feel like much when you eventually reach them. Last Thursday, I woke up sleep-deprived after working on a grant all night and found an NCBI update in my mailbox. Primer Part 2 of the genetic literacy series of ILAE Genetics Commission was now published in Epilepsia and available on PubMed. Finally, both the introductory primers of the genetic literacy series are out – Part 1 dealing with the building blocks including general concepts of epilepsy genetics and epidemiology and now Part 2 about the paradigm shifts that were introduced with the advent of massive parallel sequencing. Both publications were revised and re-written over and over again to fit the overall didactic mission of the literacy series, an effort that takes approximately 10x as long as writing a typical review. But finally, as of May 10, 2018, both Primers are now in their final shape, published and open access to the international epilepsy community. And here is just a quick overview of what this paradigm shift is really about.
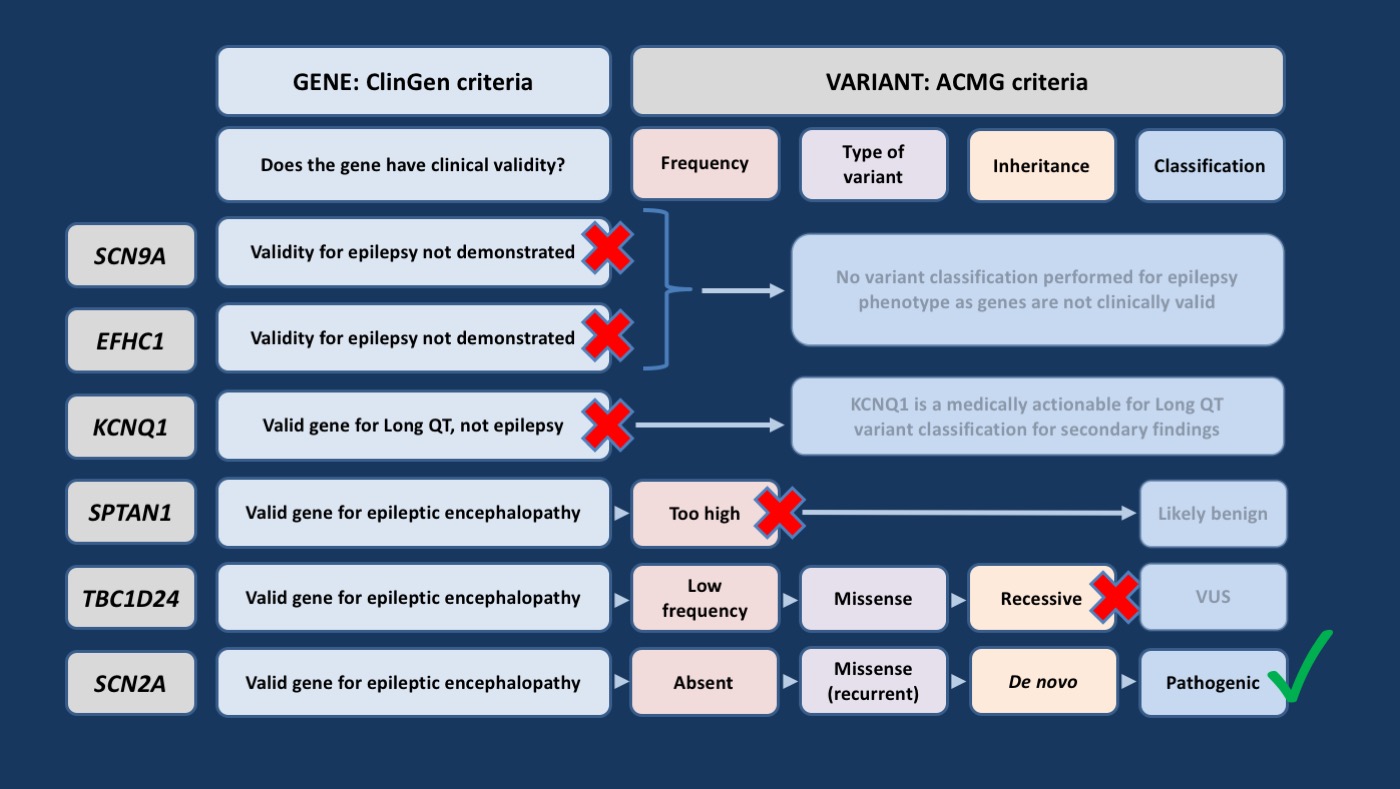
Figure 1. Example of interpretation process for determining the clinical relevance of genetic variants described in the vignette (recurrent variant refers to variants previously seen in individuals with a similar phenotype). ACMG, American College of Medical Genetics and Genomics; VUS, variant of uncertain significance [adapted from https://www.ncbi.nlm.nih.gov/pubmed/29741288]
Case-oriented. Our cases in the Genetic Literacy Series start with a case vignette – for Primer Part 2, we presented a case that is representative of scenarios in our epilepsy genetics clinic and many other neurogenetics clinics around the world. You are seeing a young boy with an unexplained epileptic encephalopathy and variants in SCN9A, EFHC1, SPTAN1, KCNQ1, SCN2A, and TBC1D24 that were identified on past genetic testing a few years ago and were considered of uncertain significance. We then guide the reader through the modern concepts of assessments of gene validity, variant pathogenicity, and clinical correlation. The figure above summarizes the detective work of variant interpretation in this patient that eventually led us to the realization that the SCN2A variant seen in this patient has acquired additional evidence since it was first reported out and is likely the cause of the patient’s disease.
A typical case. Our main reason for putting together Primer Part 2 in this format is the paradigm change that we have alluded to in the title of our manuscript. While we are still struggling to obtain genetic testing in a significant number of patients, in other patients who do undergo genetic testing, a second set of problems is emerging – the incorrect interpretation of genetic testing and reliance on outdated interpretation. Basically, results of genetic testing are not static over time. Negative and uncertain test results should ideally be reviewed and re-interpreted every 12-24 months. In fact, the “reportedly negative testing in the past” is one of the more common causes of false information that we see in clinical practice.
Summary. With the Primer Part 2, we intended to provide the clinician with a guideline to understand and question results from genetic testing to enable correct interpretation. The important thing for me to emphasize is that part of the interpretation will always be with the clinician. In the same way that we review MRIs in person, we should have a basic understanding of how genetic data is analyzed. In the same way that we don’t expect the same degree of scrutiny from the overnight radiologist who reads urgent MRIs and CTs compared to the detailed radiological discussion in a surgical case conference, we should be aware that the epilepsy-specific gene and variant interpretation will always be variable and that it will be our task to remain vigilant and skeptical to make sure that we get the best information for our patients and for the treatment choices that we are making.
Where to go from here. After reading Primer Part 2, you will hopefully be familiar with some of the current concepts that are used for gene and variant interpretation – you will understand why we are not concerned about a variant in EFHC1 in our patient as the gene lacks validity and why the SPTAN1 variant initially looks like a good candidate, but is ruled out because of the frequency of this variant in the population. You will also have been confronted with the relatively formal ClinGen vocabulary that is used in the diagnostic space. If you feel that your interest is piqued on how the process of gene and variant curation is playing out on a day-by-day basis, you might be interested in learning more about the ClinGen Epilepsy Gene Curation Expert Panel that is tasked with epilepsy gene curation and providing gene-specific information in a format that is typically used in a diagnostic setting. The clinical epilepsy world and the diagnostic world are drifting more and more apart and the ClinGen Epilepsy Gene Curation Expert Panel is currently one place where we speak both languages.



Excellent update, thanks!
Thank you!
The issue of reanalysis of ‘negative’ results is so critical and one we are struggling with in Australia as different labs have different re- reporting protocols. It is therefore so great to see this issue highlighted as it will assist ongoing constructive conversations between labs and clinicians. Thank you!
Dear Emma,
thank you for weighing in! I agree that re-analysis will remain a major issues going forward. One thing I wanted to point out is CURE’S Epilepsy Genetics Initiative (https://www.cureepilepsy.org/egi/index.html), which also aims to serve this purpose. We already had a few interesting findings coming out of EGI.
Absolutely- EGI is a fantastic initiative, very forward thinking, and one we are hoping to recruit our patients without a clear diagnosis into.