GABA/glutamate. Even though large-scale Copy Number Variation (CNV) studies in schizophrenia, autism, and intellectual disability were en vogue only a few years ago, it became quiet around these type of studies more recently. Now, a recent study in Neuron revisits CNV analysis in schizophrenia with the largest sample size to date, cancelling out much of the noise that smaller studies had to fight with. Find out how the authors managed to demonstrate a contribution of the GABAergic and glutamatergic system to schizophrenia by sheer numbers.
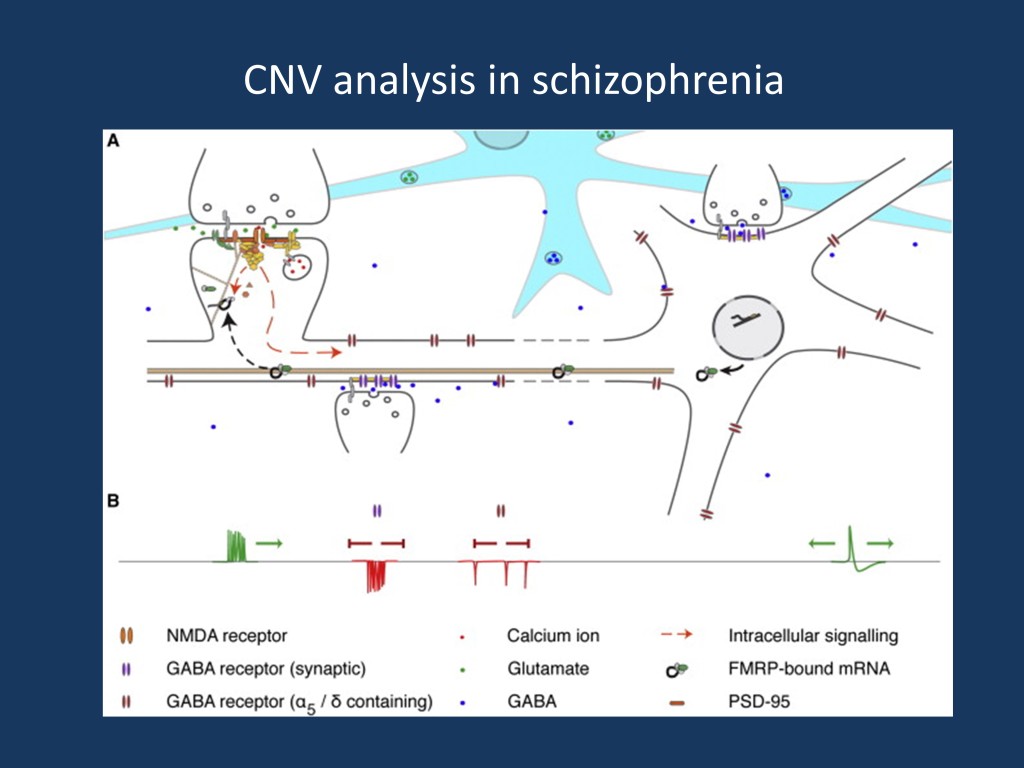
Molecular mechanisms affected in schizophrenia. The current figure is taken from the publication by Pocklington and collaborators, which is published under a Creative Commons license. The authors demonstrate through genetic studies alone that the mechanisms mentioned above are affected in schizophrenia – gene sets comprising components of the above functional compartments are affected by deletions and duplications, a finding that is highly robust on the genetic association level (Figure adapted from http://www.cell.com/neuron/abstract/S0896-6273%2815%2900372-4)
CNVs. The field of schizophrenia genetics pioneered some of our understanding of copy number variations in human disease. Slowly following the enormous success in autism, large-scale studies in schizophrenia first pinpointed genetic risk factors for neuropsychiatric disorders almost a decade ago. Ever since the publication of these impressive studies, several recurrent microdeletions are firmly established as risk factors for schizophrenia. The most prominent deletion is the 22q11.2 deletion, the microdeletion found in DiGeorge Syndrome. The initial studies robustly implicated this microdeletion in schizophrenia and also suggested other risk loci such as the 16p11.2 microdeletion. The current study by Pocklington and collaborators tried to look beyond these established loci to assess the overall contribution of rare deletions and duplications to schizophrenia.
Gene sets. The present study by Pocklington and collaborators is both impressive and unusual. The authors analyze data from 11,000 cases and 16,000 controls, making this the largest CNV association study in the field to date, which is the impressive part of this study. The unusual aspect is the strong focus on gene sets. Rather than focusing on individual candidates, the authors primarily analyze gene sets. For example, the gene set “abnormal associative learning” includes 193 genes that had been previously found to be associated with this feature through functional studies. The MGI Mammalian Phenotype Database collects the genes associated with the respective features. In this database, the term “seizures” has 783 genotypes with 1070 annotations. Working with gene sets rather than individual genes has much greater statistical power to detect differences between cases and controls. For their current study, the authors looked at 134 gene sets that were relevant to CNS function.
Permutations. One of the main analyses in their publication is a permutation analysis. The authors asked the question how many CNS gene sets showed significant enrichment in cases with schizophrenia compared to controls. The authors used a permutation analysis where they compared the observed number of enriched gene sets with the number of gene sets that they observe if they randomly mix up CNVs in cases and controls. They come to the conclusion that their observation of 23 enriched gene sets when analyzing both deletions and duplications is much more than they would expect by chance (1.3 gene sets). But which gene sets are those?
GABA/glutamate. Genes related to the NMDA receptor network and GABA-A receptor signaling are amongst the most robust finding in the study. These results even hold true if the authors look at deletions and duplications separately and remove known schizophrenia loci such as the 22q deletion. This is particularly important since some associated regions such as the 15q duplication carry multiple GABA-A receptor genes, potentially biasing the analysis. However, given the enormous size of their study, these results hold true even when these loci are disregarded. Interestingly, only CNS-related gene sets are enriched. Again, this is likely due to the size of their study.
Candidate genes. Even though their study largely looks at gene sets, the authors also focus on the most relevant candidates within the microdeletions and microduplications. Outside of the known, recurrent microdeletions, there is a strong enrichment for CNVs spanning SCL1A1, GABRD, DLG2, DLGAP1, and SHANK2. Also, PCLO and NSF are associated genes coding for proteins in the presynaptic release machinery. Other candidate genes overlap with known recessive Mendelian disorders, including AP4E1, CLN3, ISPD, and POMK.
This is what you need to know. Pocklington and collaborators present the largest CNV association study in neurological disorders so far, demonstrating that sample sizes above 10,000 cases and controls combined with an analysis of gene sets allow us to look deeper into the genetic architecture of a highly genetically heterogeneous disease. Their study convincingly demonstrates the role of both glutamatergic and GABAergic dysfunction in schizophrenia and may serve as a model for what we can achieve in epilepsy genetics in the next few years.


