Exome rounds. Interpreting genetic variants is one of the main challenges in genomic medicine. Many people have perceived barriers to starting some of the variant analysis themselves, given that there is the widespread notion that this requires expert bioinformatics knowledge. However, this is somewhat a concept of the past. There are some beautiful and simple tools online that you can use for free. Here are my favorite web-based tools for variant interpretation.
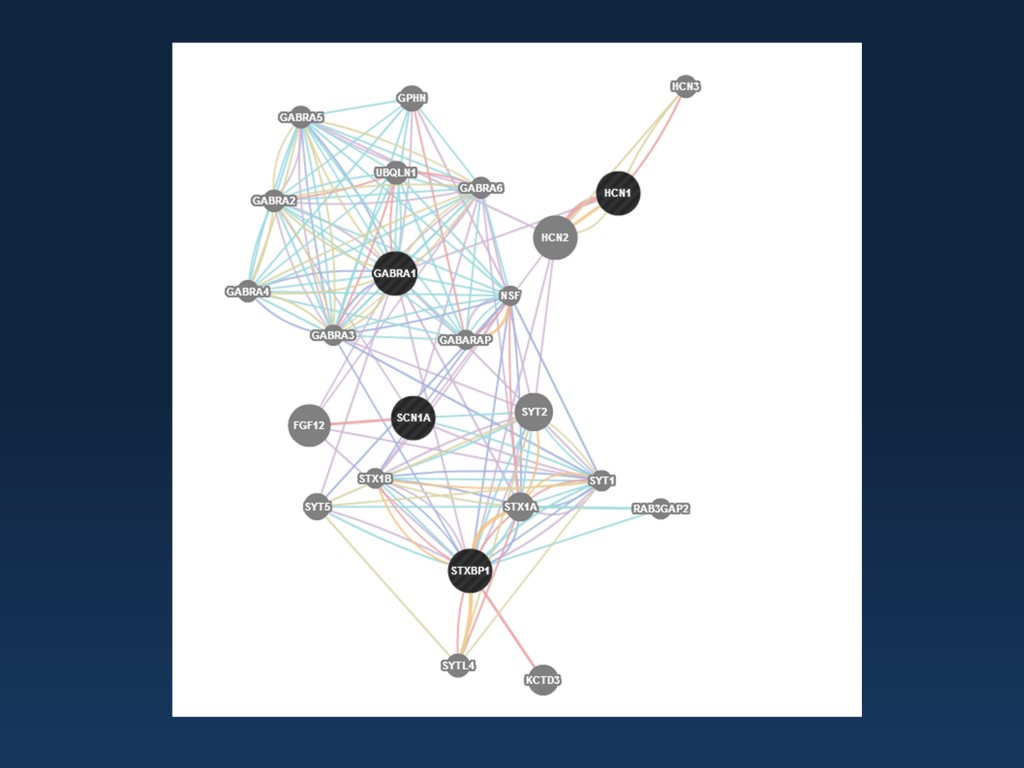
Screenshot of genemania looking at the connection between SCN1A, STXBP1, HCN1, and GABRA1. As recommended by the authors, I am citing the initial publication for this web-based tool as a reference: The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function; Warde-Farley et al., Nucleic Acids Res. 2010 Jul;38(Web Server issue):W214-20. doi: 10.1093/nar/gkq537)
Disclaimer. The tools listed in this blog will not allow you to perform a complete exome analysis. This is really beyond the scope of this blog post. In contrast, I wanted to list a few things that will help you with one of the most common problems when it comes to clinical epilepsy genetics: you have variant list, either generated in a diagnostic or research context and you simply don’t know how to interpret the findings. These tools should help you.
wANNOVAR. I have written a separate post about the web-based version of annovar, as I was really amazed by the efficiency and the speed of this program. Annovar helps with variant annotation and tells you about the frequency of a given variant in various databases including ExAC. Also, it uses a wide range of functional annotation including the CADD score. Taken together, you get a complete annotation of your variant list that will help you prioritize candidate genes.
Mutalyzer. At first, the name of this program may sound like the title of a new Lordi song, but this program has little to do with Finnish Hard Rock. Mutalyzer accomplishes the amazing feat of converting cDNA positions back to genomic positions and vice versa. Sometimes, all you get is a cDNA positions. Until I discovered Mutalyzer, this was almost an impossible task for me. Now all it takes is a single click.
RVIS. Looking at genic intolerance is a good way to prioritize candidate genes for neurodevelopmental conditions and we have features this tool in a previous post. RVIS has solved the long-standing questions of how we can safely look at exomes without constantly been confronted with MUCs, GPR98, and Titin. For any novel gene, RVIS is some kind of a litmus test and if the value is higher than the 80-90% percentile, you will get very, very skeptical. A new addition: the scores have been re-calculated for the ExAC dataset, an RVIS on steroids, so to speak.
MGI. Is there a mouse model for the gene that I’m interested in? Check MGI, the website of Mouse Genome Informatics provided by the Jackson labs. There is a nice tool to look for Human/Mouse gene connection (link). However, be aware that this tool sometimes adds some genes that you didn’t expect. For example, if you look for SCN2A, it will also give you SCN9A as a homologous gene in the mouse.
Genemania. When it comes to visualizing possible interaction partners, I had to make a decision between genemania and the STRING database. Both tools give you a quick overview about the genes and proteins that interact with your candidates and help you understand the functional pathways.
Anything else? Is there any other fast, web-based tool that I’m missing that doesn’t require more than a laptop and an internet connection? Please let me know and I’d be happy to add it to this list.


